Plants recognise pathogenic and beneficial microorganisms
In collaboration with national and international experts, researchers from Aarhus University have revealed new fundamental features of biomolecular interactions that enable plants to identify and respond appropriately to microorganisms. The new results provide a better understanding of the mechanisms governing the ability of plants to interact with beneficial microorganisms while being resistant to pathogenic bacteria and fungi. This could have implications for future sustainable agriculture, where useful microorganisms are increasingly sought to replace pesticides.
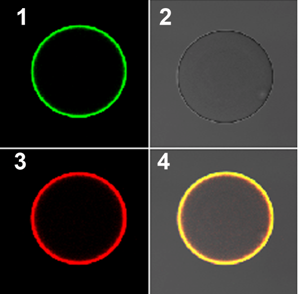

Plant roots are surrounded by thousands of bacteria and fungi living in the soil and on the root surface. To survive in this diverse environment, plants employ sophisticated detection systems to distinguish pathogenic microorganisms from beneficial microorganisms.
Here the so-called chitin molecules from microorganisms, along with modified versions, play an important role as they are detected by the plant surveillance system. Legumes, for example, build a defence against pathogenic microorganisms in response to simple chitin molecules.
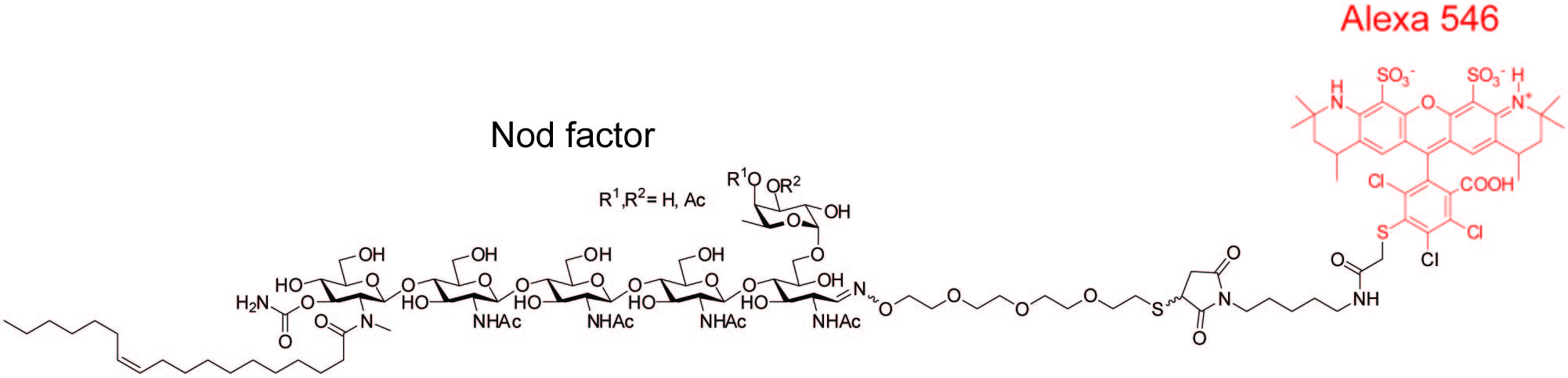
However, when the plant detects a specific modified chitin molecule (called a Nod factor) that is secreted from the rhizobium soil bacteria, formation of new organs in the form of "root nodules" occurs. Rhizobium bacteria are allowed to enter and colonise in these symbiotic organs, and they ultimately produce nitrogen for the plant.
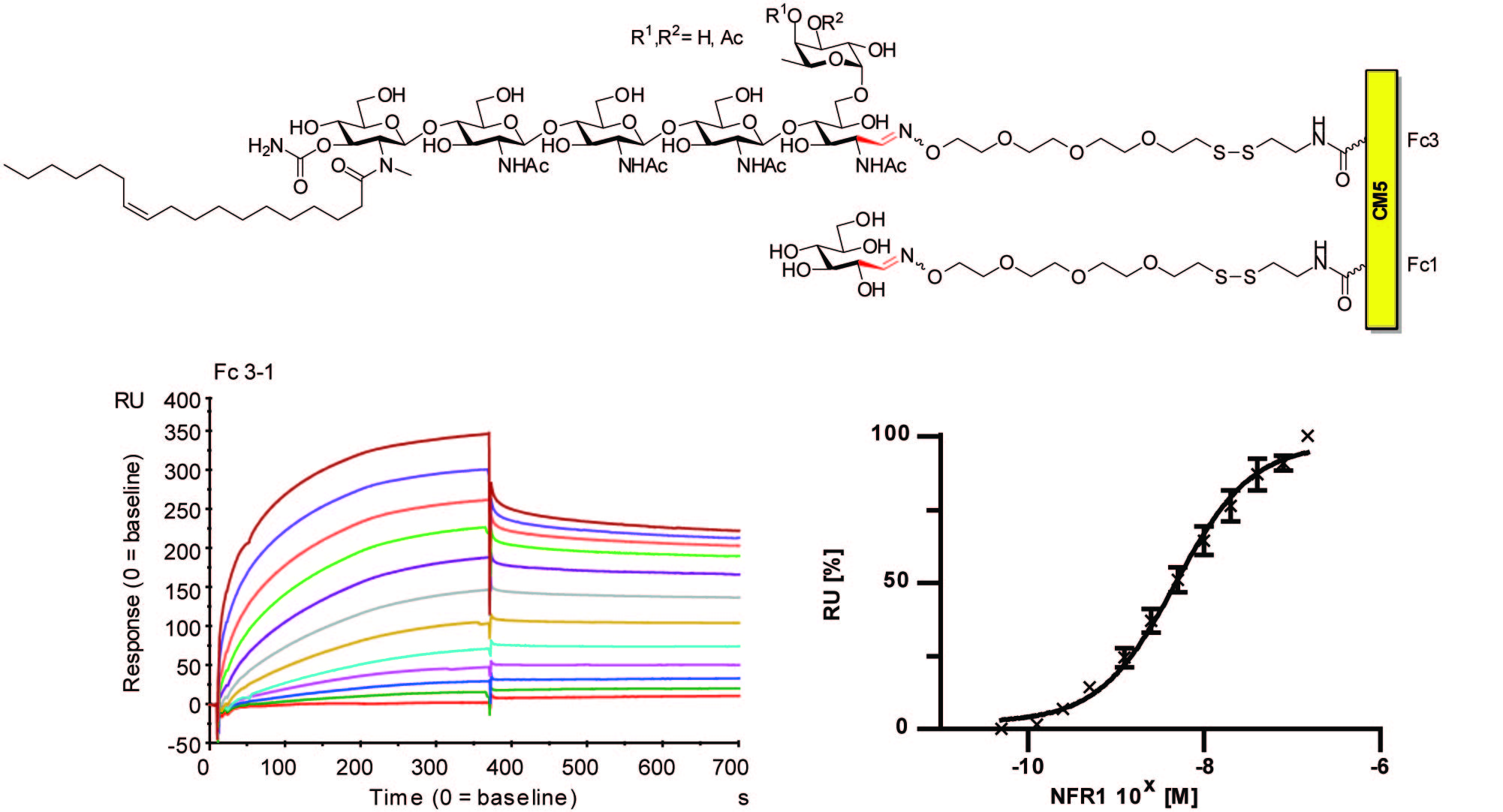
The plants’ detection of ligands – such as chitin and modified Nod factors – takes place through protein receptors that are localised on the surface of cells. Research at the Centre for Carbohydrate Recognition and Signalling (CARB) has now shown that ligand recognition through direct Nod factor binding is a key step in the receptor-mediated signal transduction that leads to root nodule development in legumes.
High-affinity binding was observed in the nano-molar range, comparable to the biologically relevant concentrations where Nod factor has in vivo activity. In contrast to this, simple chitin molecules bind to the receptors with low affinity. Structure-dependent ligand specificity and ligands binding affinities at different receptors may therefore determine which response mechanism is activated in plants exposed to different microbes or microbial communities in the environment.
Interdisciplinary approaches combining advanced biochemistry, chemoselective chemistry and microbial genetics made it possible to investigate the molecular mechanisms involved in distinguishing between Nod factor molecules secreted from rhizobia and chitin secreted by pathogenic microorganisms.
The challenging task of purifying plant receptor proteins, which are present in very low amounts in roots of the model legume Lotus japonicus, was successfully accomplished by expressing the receptors in heterologeous plant-based systems and purifying them from membrane fractions.
Another challenge was the establishment of binding assays with the carbohydrate ligands. Nod factor labelling and Nod factor immobilisation facilitated this, following application of chemoselective chemistry.
The researchers behind the results that have just been published in the international journal PNAS are affiliated with the Danish National Research Foundation’s Centre for Carbohydrate Recognition and Signalling at the Department of Molecular Biology and Genetics, Aarhus University (Denmark), Department of Chemistry, University of Copenhagen (Denmark) and Department of Microbiology and Immunology, University of Otago (New Zealand).
Link to the scientific article in PNAS: Legume receptors perceive the rhizobial lipochitin oligosaccharide signal molecules by direct binding
Angelique Broghammera,b, Lene Krusella,b, Mickaël Blaisea,b, Jørgen Sauera,c, John T. Sullivana,d, Nicolai Maolanona,c,
Maria Vinthera,b, Andrea Lorentzene, Esben B. Madsena,b, Knud J. Jensena,c, Peter Roepstorffe, Søren Thirupa,b, Clive W. Ronsona,d, Mikkel B. Thygesena,c, and Jens Stougaarda,b
aCentre for Carbohydrate Recognition and Signalling;
bDepartment of Molecular Biology and Genetics, Aarhus University, DK-8000 Aarhus, Denmark;
cDepartment of Chemistry, Faculty of Science, University of Copenhagen, DK-1871 Frederiksberg C, Denmark;
dDepartment of Microbiology and Immunology, University of Otago, Dunedin 9054, New Zealand;
eDepartment of Biochemistry and Molecular Biology, University of Southern Denmark, DK-5230 Odense M, Denmark
Contact
Professor Jens Stougaard
Director of the Danish National Research Foundation’s Centre for Carbohydrate Recognition and Signalling (CARB)
Department of Molecular Biology and Genetics, Aarhus University, Denmark
stougaard@mb.au.dk - +45 6020 2649
Text: Jens Stougaard and Lisbeth Heilesen
Translation from English into Danish: Lisbeth Heilesen
1 November 2012
