How cells in the immune system eliminate microorganisms and diseased tissue
Danish researchers have determined in atomic detail how an important mechanism in the immune system works. These results could form the basis for improved pharmaceuticals.

The complement system is an important part of immune defence and is found in all higher organisms, from fish to humans. The complement system contributes to eliminating both damaging microorganisms and diseased tissue.
When the microorganisms or diseased tissue are recognised, the complement system forms small proteins that attract and activate white blood cells. The complement system also marks out the unwanted cells by binding complement proteins to their surface. This makes it possible for immune cells called macrophages to digest the unwanted cells by means of two protein receptors called CR3 and CR4.
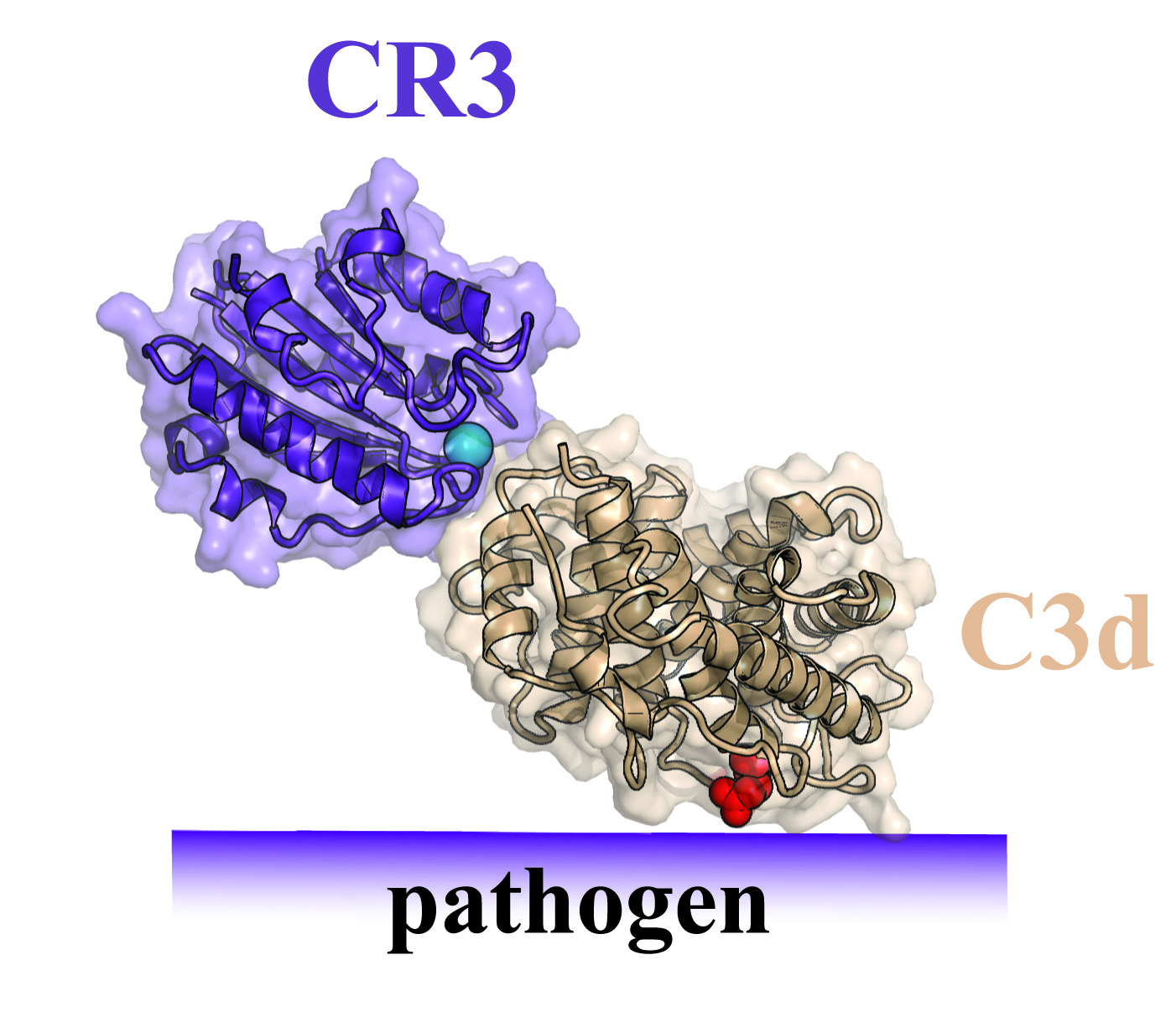
The function of these receptors has been the object of studies for many years, but the research team has now demonstrated using X-ray crystallography how the CR3 receptor recognises a particular complement protein called C3. The researchers have actually succeeded in determining the three-dimensional structure of the complex between a fragment of the CR3 receptor and a fragment of the C3 protein, which becomes bound to the surface of pathogenic organisms or damaged tissue.
“Being able to map important molecular interactions between complement proteins and immune cell receptors means we can show the way towards improving drugs for treating medical conditions where the complement system leads to unwanted tissue damage. There’s a growing interest in the way the complement system contributes to diseases with a low – but chronic – activation of processes that cause inflammation,” says Professor Gregers Rom Andersen, Department of Molecular Biology and Genetics. “Better control of these processes no doubt has the potential to reduce conditions such as the late complications of diabetes. Our discovery does not in itself show how such a control could be created, but it provides an opportunity for a more systematic approach to developing relevant pharmaceuticals,” continues Professor Andersen.
“Regulating the function of CR3 by means of pharmaceuticals is no easy task, as this receptor has a number of functions. We now know very precisely which part of the C3 protein is in contact with CR3, and this gives us better opportunities to develop ways of controlling the activity of the receptor,” he concludes.
Professor Thomas Vorup-Jensen, Department of Biomedicine, is also very satisfied with the new results, and he is convinced that his collaboration with Professor Andersen’s group will pave the way towards an even deeper understanding of these fundamental reactions in the immune system.
- “Our new results have clarified the intimate molecular mechanisms in a key part of the immune system, which was discovered more than thirty years ago by Timothy A. Springer at Harvard University. In terms of pure science, it’s extremely satisfying to finally be able to visualise the function of molecules in the immune system by means of X-ray crystallography,” he concludes.
The process was a special experience for PhD student Goran Bajic, Department of Molecular Biology and Genetics.
- “It’s madly exciting to be the person to see for the first time how millions of years of evolution have made it possible for our immune system to gain control over pathogenic microorganisms and damaged tissue. This is a rare moment, putting together all biochemical data like the pieces of a huge, surprising jigsaw puzzle,” he says. Goran Bajic carried out a major part of the experiments.
The results have just been published in the international journal PNAS (Proceedings of the National Academy of Sciences).
Link to the scientific article in PNAS: Structural insight on the recognition of surface-bound opsonins by the integrin I domain of complement receptor 3
Goran Bajica, Laure Yatimea, Robert B. Simb, Thomas Vorup-Jensenc, and Gregers R. Andersena
Departments of aMolecular Biology and Genetics and cBiomedicine, Aarhus University, DK-8000 Aarhus, Denmark; and
bDepartment of Pharmacology, University of Oxford, Oxford OX1 3QT, United Kingdom
Professor Thomas Vorup-Jensen, Department of Biomedicine, Aarhus University, has studied the function of CR3 and CR4 since he worked as a postdoctoral fellow in Professor Timothy A. Springer’s laboratory at Harvard University. Professor Gregers Rom Andersen, Department of Molecular Biology and Genetics, Aarhus University, has for many years used X-ray crystallography to study complement proteins.
PhD student Goran Bajic carried out the experimental work under the supervision of Professor Andersen and Professor Vorup-Jensen. Data produced by Professor Vorup-Jensen was first analysed with the help of Dr Robert B. Sim, University of Oxford. This analysis revealed that the two receptors – CR3 and CR4 – recognise the C3 complement protein in completely different ways. By using gene splicing, Goran Bajic then produced fragments of the C3 and CR3 proteins, with the assistance of Senior Adviser Laure Yatime, and formed crystals containing the two proteins. These were then clarified using X-radiation, and the three-dimensional structure of the C3-CR3 complex could finally be produced.
In connection with a grant from the Lundbeck Foundation to set up the Lundbeck Foundation Nanomedicine Centre for Individualized Management of Tissue Damage and Regeneration (LUNA), the two professors entered into collaboration to study the function of CR3 and CR4. This work has now resulted in the researchers being able to show how CR3 works in atomic details.
More information
Professor Gregers Rom Andersen
Department of Molecular Biology and Genetics
Aarhus University
gra@mb.au.dk
+45 8715 5507
or
Professor Thomas Vorup-Jensen
Department of Biomedicine
Aarhus University
vorup-jensen@microbiology.au.dk
+45 2148 9781
