New insights into how the CRISPR immune system evolved
With new insights into how the genetic tool CRISPR – which allows direct editing of our genes – evolved and adapted, we are now one step closer to understanding the basis of the constant struggle for survival that takes place in nature. The results can be used in future biotechnologies.

In 2020, the Nobel Prize in Chemistry goes to Emmanuelle Charpentier and Jennifer A. Doudna for their discoveries of the molecular mechanism behind CRISPR-Cas and the use of the technology as a genetic tool. Although CRISPR-Cas has found many uses in biotechnology and medicine, it originates in nature, where it functions as a microbial immune system.
Just as our immune system remembers the pathogens we have been exposed to earlier in life, CRISPR-Cas provides microorganisms with an ability to respond quickly to viruses they have previously encountered by storing a small amount of the viral DNA in their own genome.
CRISPR-Cas is found naturally in most bacteria as well as the so-called archaea. When examining the origin of life on Earth, archaea are particularly interesting, as they form a kind of "missing link" between bacteria and the cells of higher eukaryotes like our own. Studies of these organisms can therefore provide us with important insights into how the CRISPR-Cas immune system has evolved over hundreds of millions of years.
New results reveal why toxins are present among CRISPR-Cas genes
New research results from researchers at the Department of Molecular Biology and Genetics, Aarhus University – obtained in close collaboration with leading researchers from the University of Copenhagen and Old Dominion University in Virginia, USA, and published in two articles in leading, international journals – now shed new light on how CRISPR-Cas emerged early during the development of life on Earth, as well as how this immune system is constantly adapting to new challenges.
The research group from Aarhus – led by Associate Professor Ditlev E. Brodersen – has discovered how a part of CRISPR-Cas that is responsible for incorporating foreign, viral DNA into the microorganism's own genome has originated from another, very common type of genes in bacteria and archaea that surprisingly encode toxins.
The new knowledge therefore provides insights into an evolutionary process in which the toxin genes were present early during the development of life, and over time were integrated and adapted as part of the CRISPR-Cas modules that many microorganisms possess to this day. For the first time, we have an answer to a question that has puzzled researchers for a long time, namely why toxin genes exist among the CRISPR-Cas genes.
- "This understanding of how certain proteins are 'recycled' in several different situations, is enormously useful for researchers," explains Ditlev Brodersen, "because when we understand the entire repertoire of functions that certain proteins possess, it opens up for the possibility of using them as specific tools in genetic engineering. For example, it might be possible to get disease-causing bacteria to direct their CRISPR-Cas systems towards themselves and thus avoid infection."
A constant battle between microorganisms and viruses
In another article, published in the renowned journal, Nature Communications, the researchers describe new findings that provide insights into the constant battle between microorganisms and the viruses that represent their worst enemies.
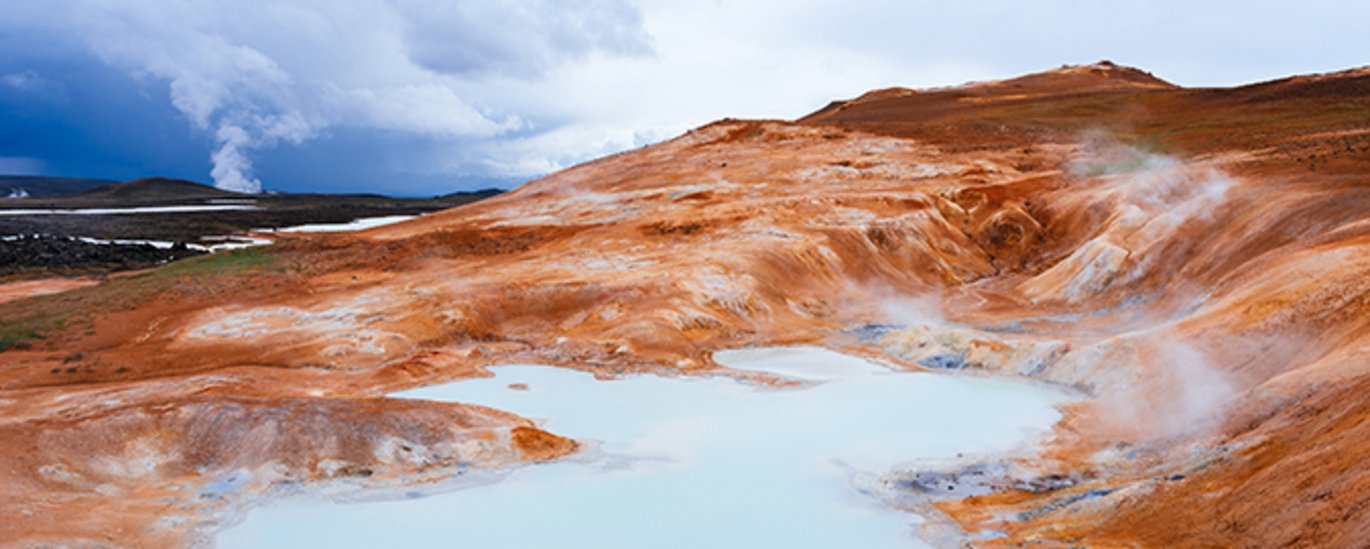
In a boiling mud puddle on Iceland lives a very special organism, an archaea called Sulfolobus islandicus, which for millions of years has adapted to life in this place, that with a constant temperature of 80-100°C and an acidity corresponding to stomach acid constitutes one of the most inhospitable places on earth.
But even though Sulfolobus has chosen a very unattractive place to live, it still encounters resistance, not the least from small, rod-shaped DNA viruses that constantly poke holes in the cells and shoot their foreign DNA into them, causing Sulfolobus to explode in a wealth of new virus particles. To avoid this fate, Sulfolobus has developed a CRISPR-Cas defence, by which it has stored small parts of the viral DNA in its own genome to be able to withstand these attacks.
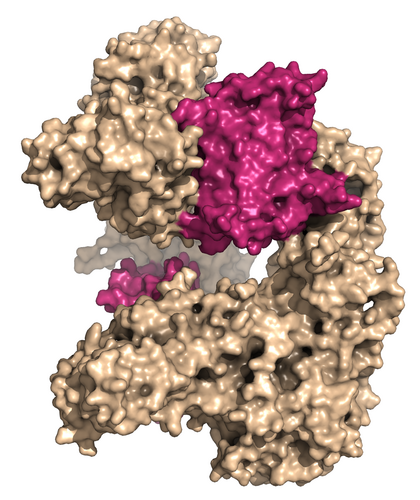
The new results show in great detail how two anti-CRISPR proteins (pink) lock two CRISPR-Cas proteins (beige) together and thus prevent them from destroying the viral DNA (D. E. Brodersen).
Anti-CRISPR – Upsetting the applecart
But in the constantly escalating battle between life and death, the virus has developed a countermeasure: It has managed to cope by producing a small weapon, an anti-CRISPR protein that, like upsetting the applecart, blocks the CRISPR-Cas response in Sulfolobus.
The new results from Ditlev E. Brodersen's group at Aarhus University – generated in close collaboration with Associate Professor Xu Peng from the Department of Biology, University of Copenhagen – now for the first time show how this fight takes place in the boiling pools.
The researchers have been able to visualise how the anti-CRISPR protein binds strongly to the largest protein of the CRISPR-Cas system, thereby directly preventing it from destroying the viral DNA. In this way, the virus bypasses – at least for some time – being beated off by CRISPR-Cas. The new results give scientists insights into the arms race that is constantly taking place in nature, and how the evolution of life is in fact a constant struggle for survival.
- "We now know the details of how the anti-CRISPR protein can block the CRISPR-Cas immune system, so the question is what will be the next move in this arms race," says Ditlev Brodersen. "Perhaps the microbes will begin to form anti-anti-CRISPR proteins, a third type of protein that can prevent the anti-CRISPR protein from working, but we have yet to find these in Sulfolobus archaea. So right now the ball is back on Sulfolobus' half of the field", says Ditlev Brodersen, "and the cold war is always warm in the boiling pool".
Link to the article in Structure:
"Structural Basis for Toxin Inhibition in the VapXD Toxin-Antitoxin System"
Marie B. Bertelsen, Meriem Senissar, Maja H. Nielsen, Ashley L. Molinaro, Dayle A. Daines, Ditlev E. Brodersen
DOI: https://doi.org/10.1016/j.str.2020.10.002
Link to the article in Nature Communications:
Structural basis for inhibition of an archaeal CRISPR–Cas type I-D large subunit by an anti-CRISPR protein
M. Cemre Manav, Lan B. Van, Jinzhong Lin, Anders Fuglsang, Xu Peng & Ditlev E. Brodersen
DOI:https://doi.org/10.1038/s41467-020-19847-x
For further information, please contact
Associate Professor Ditlev E. Brodersen
Department of Molecular Biology and Genetics
Aarhus University, Denmark
Tel. +45 21669001, email deb@mbg.au.dk
