New long-read sequencing technology reveals the surprising complexity of circular RNAs
A paper from Jørgen Kjems and colleagues published in Nature Communications presents a new method to determine the full-length sequence and isoform variation of a class of RNAs called circular RNAs. The study is conducted in human and mouse brains and shows that many circRNAs exist in a large number of different version and that a special class of exons, previously linked to autism, are enriched in circRNAs.


When cells read the information encoded in our genes, they do so by creating an RNA copy of the individual gene. RNA molecules are processed in a myriad ways, creating the complexity in gene expression seen in higher organisms. The average gene produces 7 different isoforms, largely through alternative splicing, and this process is particularly active in neurons. Circular RNAs (circRNAs) are a group of highly stable RNAs are enriched in neurons but their entire composition remain unexplored due to their sequence overlap with conventional mRNA.
One of the challenges for understanding circRNA composition is that they have so far only been described using sequencing techniques that read short stretches of RNA. The method presented by Kjems and colleagues solves this problem by developing protocols where circRNAs are specifically enriched and subjected to long-read sequencing using nanopore sequencing technology.
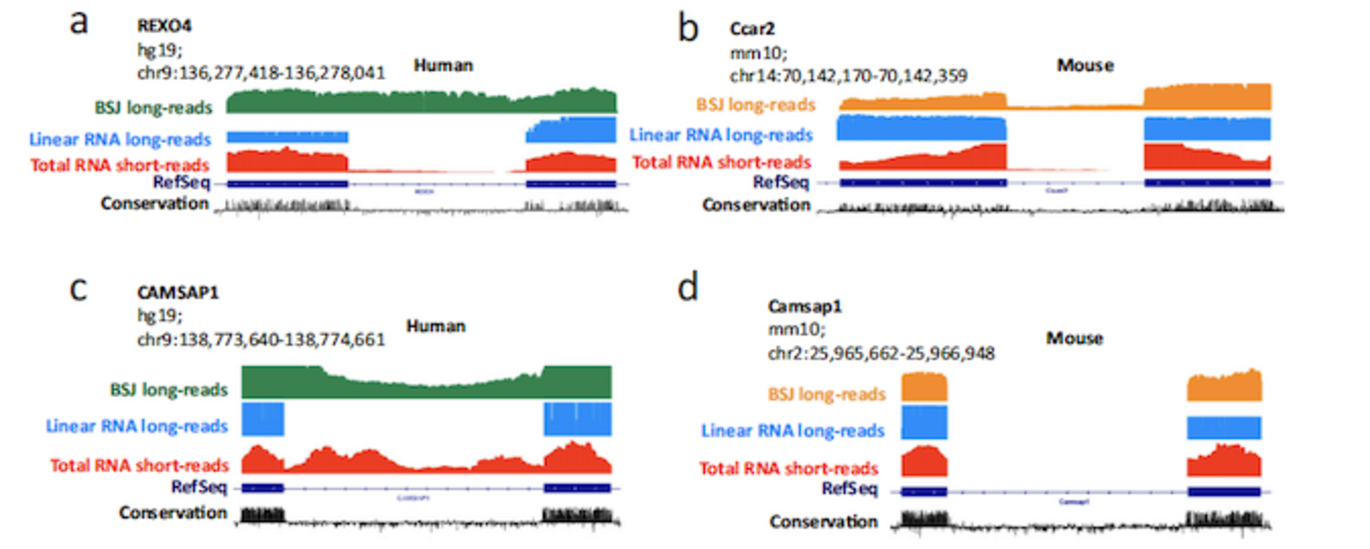
In addition to the laboratory protocol, a novel bioinformatic pipeline for the ONT long-read platform was developed by former MBG/INANO-postdoc Morten Venø - now the CEO of AU spin-out company Omiics. Taken together, this led to the surprizing finding that circRNAs exhibit more diverse splicing events than mRNA with frequent retention of introns and novel exons. Of particular interest, the authors find that a group of exons shorter than 30 nt, the so-called microexons, are preferentially found in circRNAs, rather than in linear RNAs as previously reported. Several of these microexons have been linked to autism, suggesting that the unique composition of circRNAs found in the brain could play a role in neuronal disease.
The article in Nature Communications:
Nanopore sequencing of brain-derived full-length circRNAs reveals circRNA-specific exon usage, intron retention and microexons
Karim Rahimi, Morten T. Venø, Daniel M. Dupont, Jørgen Kjems
DOI: 10.1038/s41467-021-24975-z
More information
Jørgen Kjems
iNANO/Department of Molecular Biology and Genetics, Aarhus University, Denmark
jk@mbg.au.dk – 2899 2086
Karim Rahimi
iNANO/Department of Molecular Biology and Genetics, Aarhus University, Denmark
karim@mbg.au.dk
Morten Venø
Omiics
morten.venoe@omiics.com
