Neuroplasticity underlies learning and memory formation, which allows for accumulation of knowledge. Abnormalities can be associated with various disorders including dementia and post-traumatic stress disorder (PTSD). There is an emerging view that a sparse ensemble of neurons, termed as memory engram cells, store specific memories. Memory engram cells are defined as neurons that are transiently activated and express immediate early genes (IEGs) during a learning experience and have undergone physical changes that can be reactivated upon memory recall experience. Importantly, artificial activation and silencing of engram cells result in artificial memory retrieval and deficit, respectively, and open for research approaches to unveil the mechanisms.
Revealing the molecular basis of neuroplasticity in engram cell formation is considered one of the central issues of memory research.
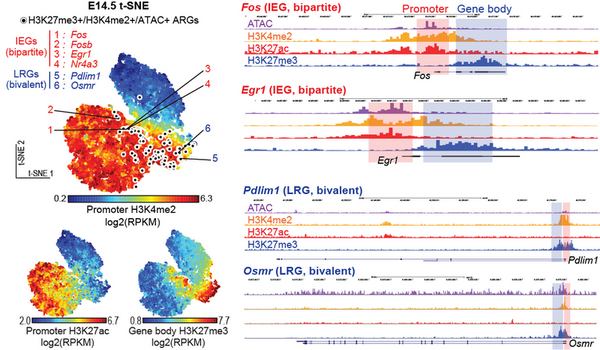
In a recent study, Taro Kitazawa and his colleagues at the Friedrich Miescher Institute for Biomedical Research (FMI) in Switzerland revealed a novel epigenetic and transcriptional mechanism regulating neuronal activity-dependent IEGs activation during somatosensory neuron maturation (Kitazawa et al., Nature Genetics, 2021, Figure).
Taro Kitazawa will now take an advantage of his research expertise and further investigate how neuronal activity-dependent gene regulation underlies memory engram cell plasticity.
In his efforts to address this issue, Taro Kitazawa will carry out epigenetic and transcriptional profiling of memory engram cells using state-of-the-art high throughput sequencing genomics technologies. Engram cells will be isolated from the mouse hippocampus or neocortical regions, which store recent (in the order of days) or remote (in the orders of weeks and months) memory, respectively.
With this approach, he will firstly reveal the molecular underpinning of engram cell state shifts upon memory transfer from the hippocampus to the neocortical regions, which has been referred to as the systems memory consolidation. Furthermore, he will also focus on the heterogeneity of memory engram cells, and aim to reveal how a subset of engram cells can be functionally more relevant as compared with other subsets of engram cells. Notably, to efficiently answer these questions, he will even develop a novel genomics technology which enables whole genome history tracing, and overcome critical limitations of existing snapshot-type sequencing technologies.