X-rays reveal the self-defence mechanisms of bacteria
A research group at Aarhus University has gained unique insight into how bacteria control the amount of toxin in their cells. The new findings can eventually lead to the development of novel forms of treatment for bacterial infections.

Many pathogenic bacteria are able to go into a dormant state by producing persister cells that are not susceptible to conventional antibiotics. This causes serious problems in the treatment of life-threatening diseases such as tuberculosis, where the presence of persister cells often leads to a resurgence of infection following medical treatment.
At the molecular level, the formation of persister cells is due to the presence of toxins that are produced by the bacteria themselves, and which enable them to enter the dormant state. During this hibernation period, the bacteria constantly regulate the amount of toxin at exactly the same level and thus maintain the dormant state.
In an article recently published in the American scientific journal Structure, the researchers at the Department of Molecular Biology and Genetics, Aarhus University, present new results that reveal the molecular details of the regulatory mechanism of toxins.
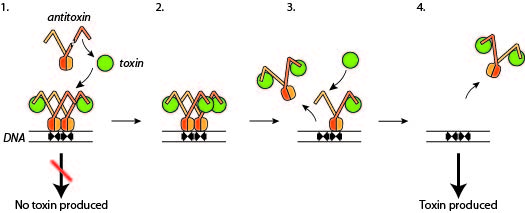
By isolating and crystallising the toxin molecules and their molecular companions – the antitoxins – and by subsequently exposing the crystals to strong X-rays, the research team (consisting of the two PhD students Andreas Bøggild and Nicholas Sofos and Associate Professor Ditlev E. Brodersen) gained unique insight into how bacteria control the amount of toxin in the cell (illustrated in the info box).
The new findings can eventually lead to the development of entirely new forms of treatment of bacterial infections that work initially by blocking toxin function and production, and subsequently by using traditional antibiotics to fight the pathogenic bacteria.
Link to the article in Structure: The crystal structure of the intact E coli RelBE toxin-antitoxin complex provides the structural basis for conditional cooperativity
Andreas Bøggild,1,3 Nicholas Sofos,1,3 Kasper R. Andersen,1,4 Ane Feddersen,1 Ashley D. Easter,2 Lori A. Passmore,2 and Ditlev E. Brodersen1
1Centre for mRNP Biogenesis and Metabolism, Department of Molecular Biology and Genetics, Aarhus University, DK-8000 Aarhus C, Denmark
2MRC Laboratory for Molecular Biology, Cambridge CB2 0QH, UK
3These authors contributed equally to this work
4Present address: Department of Biology, Massachusetts Institute of Technology, Cambridge, MA 02139, USA
More information
Associate Professor Ditlev E. Brodersen
Department of Molecular Biology and Genetics
Aarhus University
Denmark
deb@mb.au.dk – +45 2166 9001
Text: Ditlev E. Brodersen and Lisbeth Heilesen
Translation: Lisbeth Heilesen
